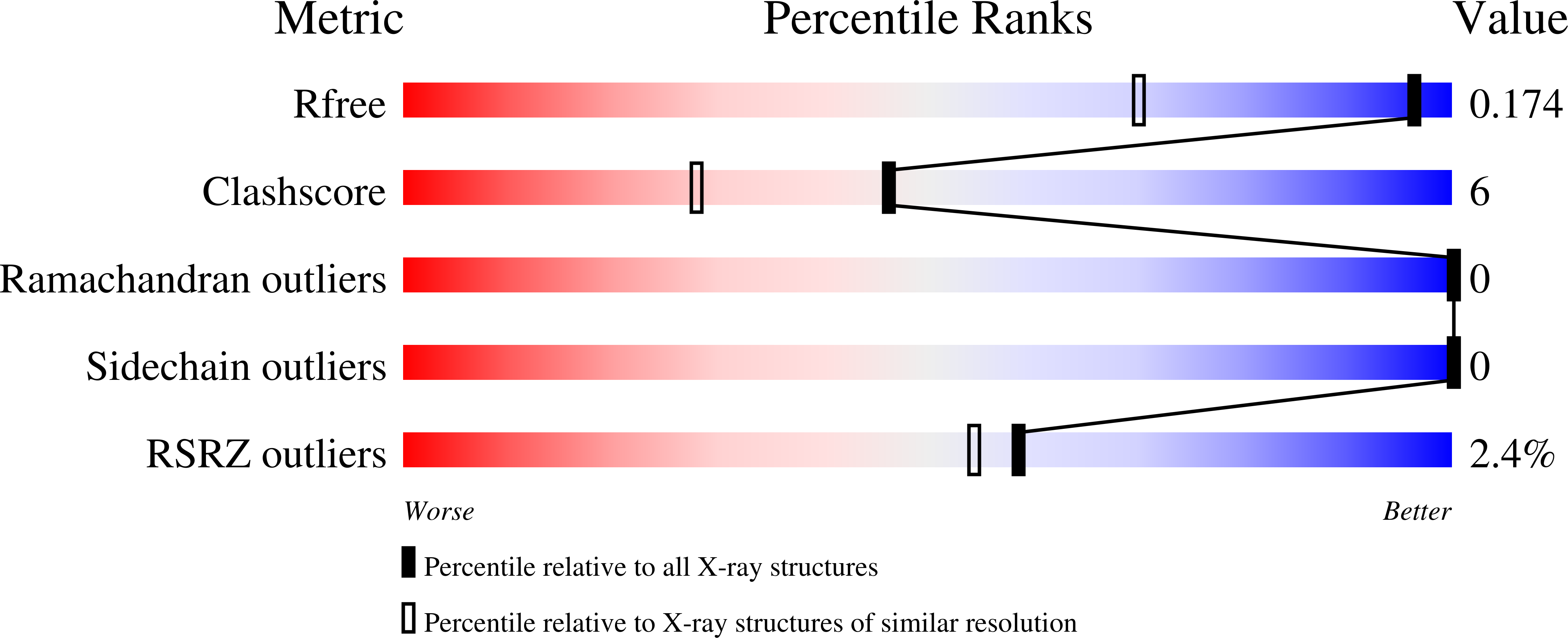
A Non-covalent KRASG12D Allele Specific Inhibitor Demonstrates Potent Inhibition of KRAS-dependent Signaling and Regression of KRASG12D-mutant Tumors
Christensen, J., Hallin, J., Bowcut, V., Calinsan, A., Briere, D., Hargis, L., Engstrom, L., Laguer, J., Medwid, J., Vanderpool, D., Lifset, E., Trinh, D., Hoffman, N., Wang, X., Lawson, J., Gunn, R., Smith, C., Thomas, N., Martinson, M., Bergstrom, A., Sullivan, F., Bouhana, K., Winski, S., He, L., Julio, F.B., Pavlicek, A., Hafing, J., Rahbaek, L., Marx, M., Olson, P.(2022) Nature